|
9/3/2023 0 Comments Cytoscape linux Visualize and analyze human-curated pathway datasets such as Reactome or KEGG. Perform advanced analysis and modeling using Cytoscape plugins Project and integrate global datasets and functional annotationsĮstablish powerful visual mappings across these data Load molecular and genetic interaction data sets in many formats For example, input a set of custom annotation terms for your proteins, create a set of confidence values for your protein-protein interactions.Ĭytoscape supports many use cases in molecular and systems biology, genomics, and proteomics: Using this feature, you can load and save arbitrary attributes on nodes, edges, and networks. Delimited text files and MS Excel™ Workbook are also supported and you can import data files, such as expression profiles or GO annotations, generated by other applications or spreadsheet programs. Most of the plugins are freely available.Ĭytoscape supports a lot of standard network and annotation file formats including: SIF (Simple Interaction Format), GML, XGMML, BioPAX, PSI-MI, SBML, OBO, and Gene Association. 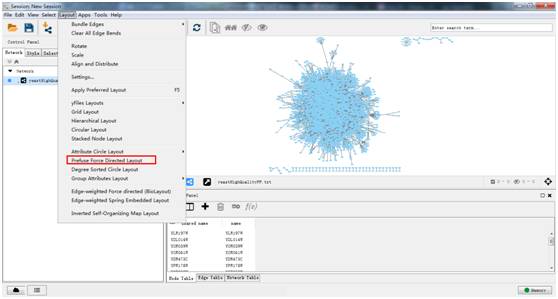
Plugins may be developed by anyone using the Cytoscape open API based on Java™ technology and plugin community development is encouraged. Plugins are available for network and molecular profiling analyses, new layouts, additional file format support, scripting, and connection with databases. Additional features are available as plugins. Cytoscape core distribution provides a basic set of features for data integration and visualization. Although Cytoscape was originally designed for biological research, now it is a general platform for complex network analysis and visualization. It has been hosted in OnWorks in order to be run online in an easiest way from one of our free Operative Systems.Cytoscape is an open source bioinformatics software platform for visualizing molecular interaction networks and biological pathways and integrating these networks with annotations, gene expression profiles and other state data. This is an application that can also be fetched from. Uses stylesheets to separate presentation from data in a rendering agnostic manner.Supports selectors for terse filtering and graph querying.Uses layouts for automatically or manually positioning nodes.Fully serializable and deserializable via JSON.Has a large suite of tests that can be run in the browser or the terminal.Supports rendering images of graphs on Node.js with Cytosnap.Some demos may not work in old browsers in order to keep the demo code simple. 
The documentation and examples are not optimized for old browsers, although the library itself is. Browsers with partial but sufficient ES5 support also work, such as IE9 and Firefox 4. 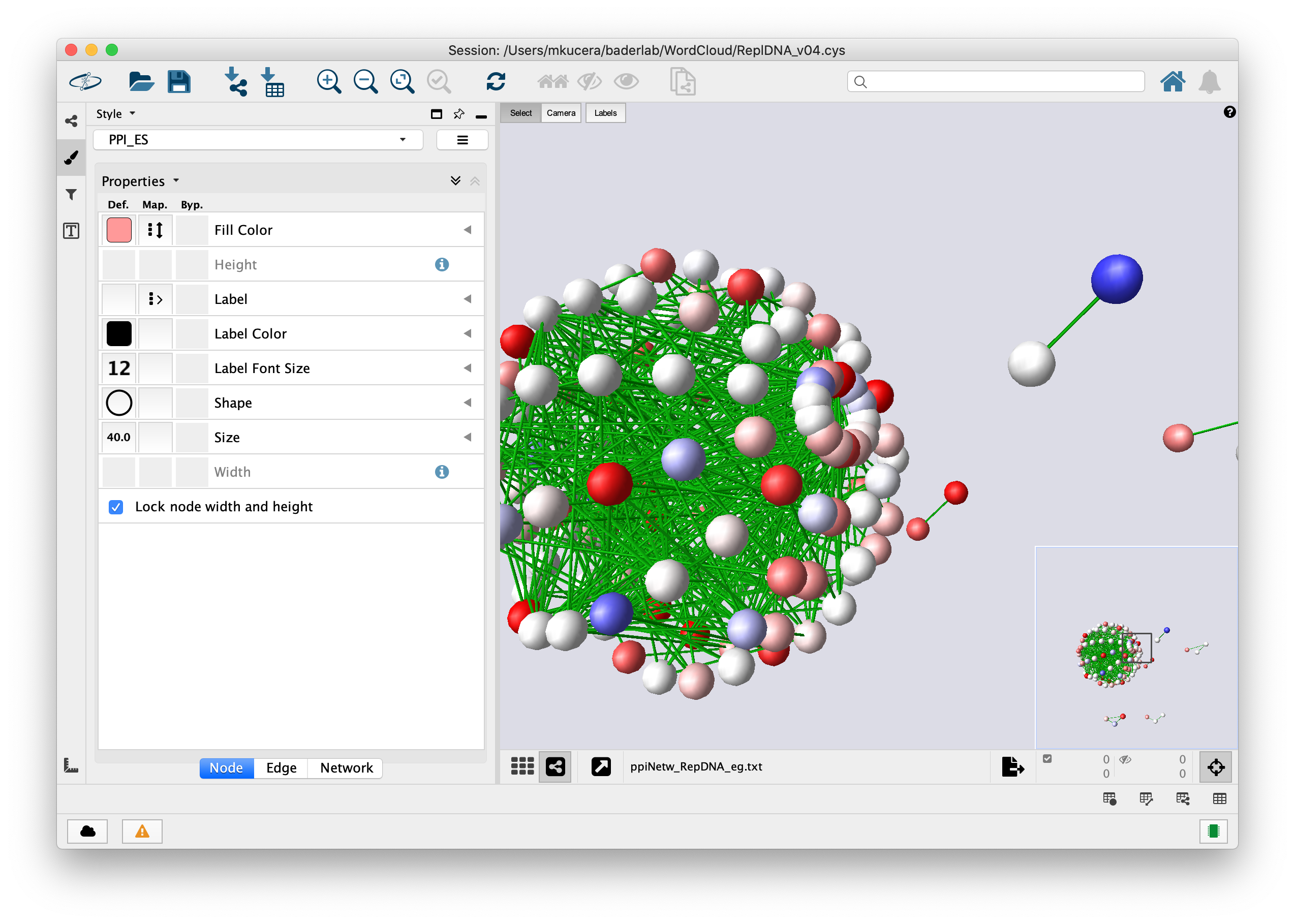
Browsers circa 2012 support ES5 fully: IE10, Chrome 23, Firefox 21, Safari 6 (caniuse). ES5 and canvas support are required, and feature detection is used for optional performance enhancements. Legacy browsers with ES5 and canvas support. Designed for users first, for both frontfacing app usecases and developer usecases. Used in commercial projects and open-source projects in production. Permissive open source license (MIT) for the core Cytoscape.js library and all first-party extensions. A fully featured graph library written in pure JS. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
